Kumar Motiram-Corral presented a poster titled “Implementing one-shot multiple-FID acquisition into homonuclear and heteronuclear NMR experiments” at SMASH 19 in Porto (Portugal).
To date, time-efficient approaches are a challenged task for spectroscopists. The goal is to obtain chemical information reducing experimental time without considerably losing of sensitivity.
Different time-efficient approaches have been described over the years. Time sharing (Parella et. al.) tactic acquires the 15N and 13C nuclei in the same spectrum in spectrometers which have a triple channel hardware configuration[1]. Non-Uniform Sampling (NUS) [2] algorithm has achieved a substantial reduction of experimental time reducing the number of t1 increments needed by multidimensional experiments. Recently, NOAH [3] (NMR by ordered Acquisition using 1H detection) has been developed by Kupče (Bruker Co.) and Claridge (University of Oxford) provides the way to get proper experiments in different spectra with the same spectral quality.
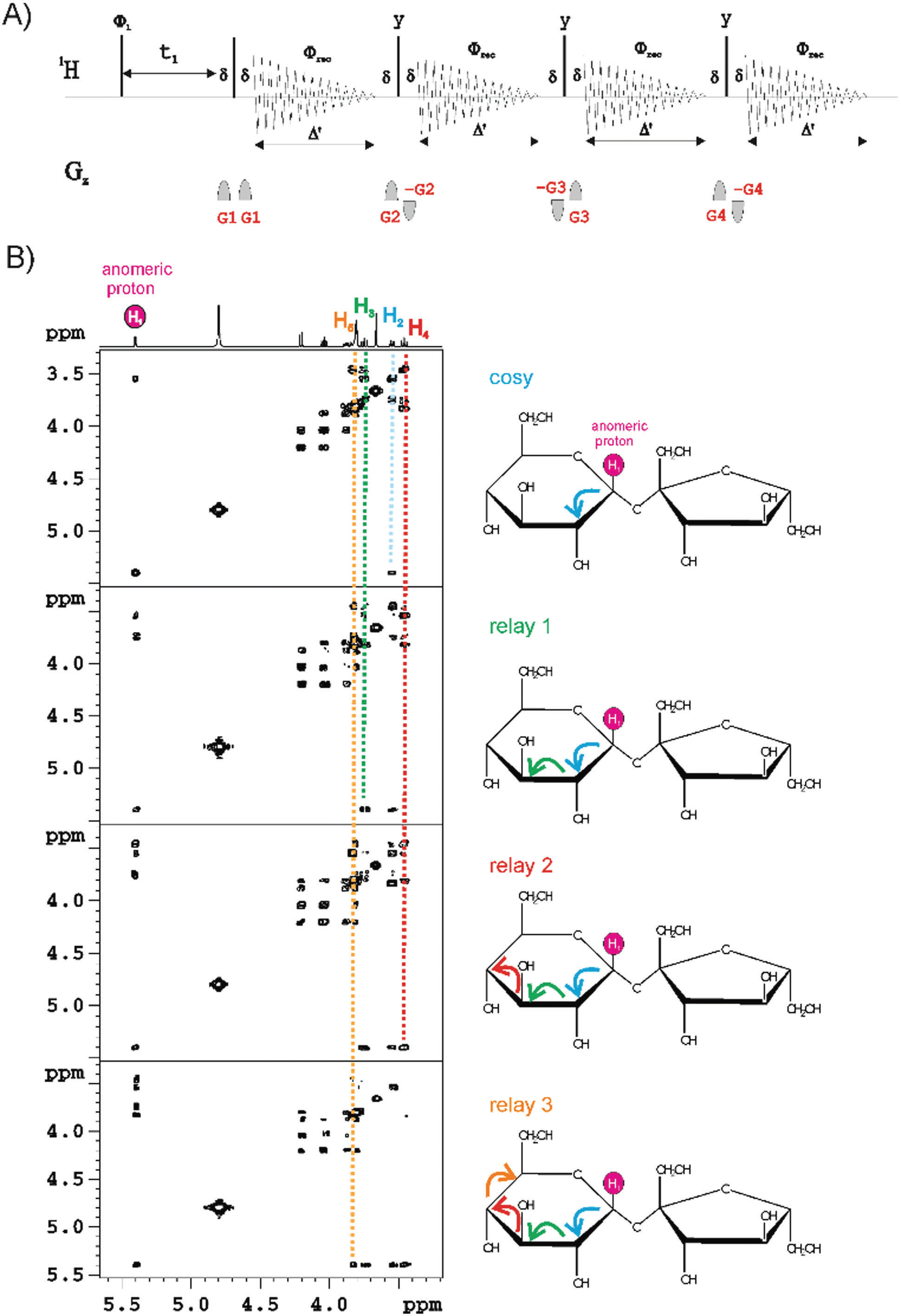
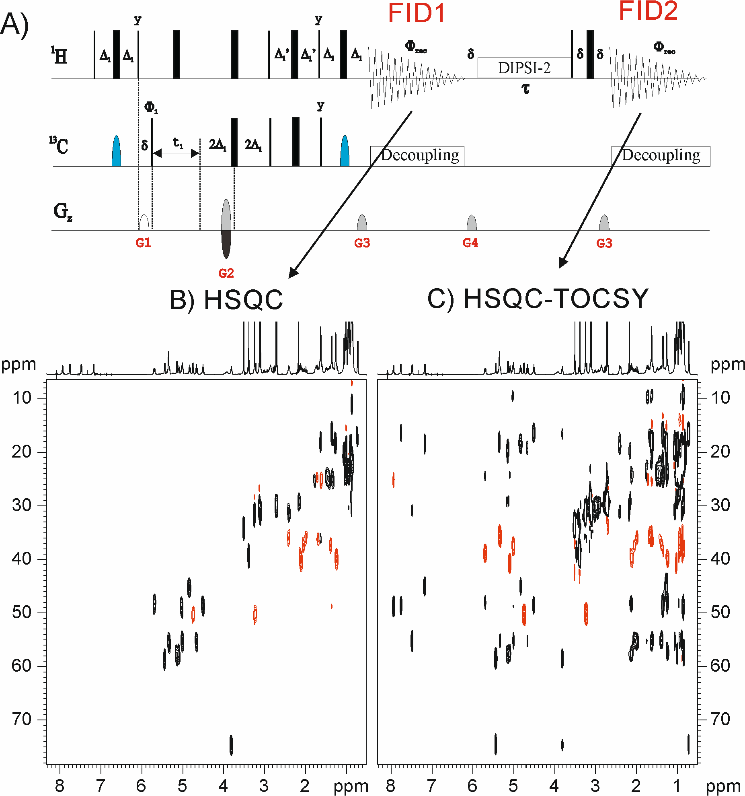
[4]MFA (Multiple FID Acquisition) consists in obtaining up to four different experiments decreasing close to 60% of time. MFA provides a new novel proof concept of COSY, TOCSY and HMBC experiments in small molecules. Actually, MFA strategy was proposed many years ago with the COCONOSY experiment [5-7], which could be collected 2D COSY and NOESY data with a single pulse scheme. [4]MFA has also been implemented in magic-angle-spinning solid-state NMR experiments devoted for biomacromolecules using standard spectrometer configuration. Despite its limitations related to the use of long acquisition of free-induction decays (FIDs) to accurately digitalize the data and the mandatory use of long phase cycles for convenient pathway selection, nowadays, the use of pulsed field gradients (PFGs) is the solution for this drawback. [4]MFA is based on the relaxation of the remaining transverse magnetization, which usually relaxes to its original magnetization, can be manipulated by an appropriate additional mixing process and recorded again to obtain a second or third NMR data provided that T2 (transverse relaxation times) are long enough. [4]Its main advantage is that each experiment is acquired in a different display. [4]MFA is a powerful experiment for the sequential structural assignment of a whole spin system without ambiguities. This method is also useful for selective experiments as SE-TOCSY.
References:
- Nolis, P., Pérez, M., & Parella, T. (2006). Time-sharing evolution and sensitivity enhancements in 2D HSQC-TOCSY and HSQMBC experiments. Magnetic Resonance in Chemistry, 44, 11, 1031-1036, 2006
- K. Kazimierczuk and V. Y. Orekhov , Angew. Chem., Int. Ed., 2011, 50 , 5556 -5559
- Kupče, E., & Claridge, T. D. W. (2018). Molecular structure from a single NMR supersequence. Chemical Communications, 54, 7139-7142, 2018.
- Motiram-Corral, K., Pérez-Trujillo, M., Nolis, P., & Parella, T. (2018). Implementing one-shot multiple-FID acquisition into homonuclear and heteronuclear NMR experiments. Chemical Communications, 54(96), 13507–13510, 2018.
- A. Z. Gurevich , I. L. Barsukov , A. S. Arseniev and V. F. Bystrov , J. Magn. Reson., 56, 471 -478, 1984.
- C. A. G. Haasnoot , F. J. M. van de Ven and C. W. Hilbers , J. Magn. Reson., 56 , 343 -349, 1984.
- J. Cavanagh and M. Rance , J. Magn. Reson., 14 , 408 -414, 1990.