“Simultaneous 1H and 13C NMR enantiodifferentiation from highly-resolved pure shift HSQC spectra” by Miriam Pérez-Trujillo, Laura Castañar, Eva Monteagudo, Lars T. Kuhn, Pau Nolis, Albert Virgili, R. Thomas Williamson and Teodor Parella. Chemical Communications 50:10214-10217 (2014). DOI: 10.1039/C4CC04077E
“Simultaneous 1H and 13C NMR enantiodifferentiation from highly-resolved pure shift HSQC spectra” by Miriam Pérez-Trujillo, Laura Castañar, Eva Monteagudo, Lars T. Kuhn, Pau Nolis, Albert Virgili, R. Thomas Williamson and Teodor Parella. Chemical Communications 50:10214-10217 (2014). DOI: 10.1039/C4CC04077E
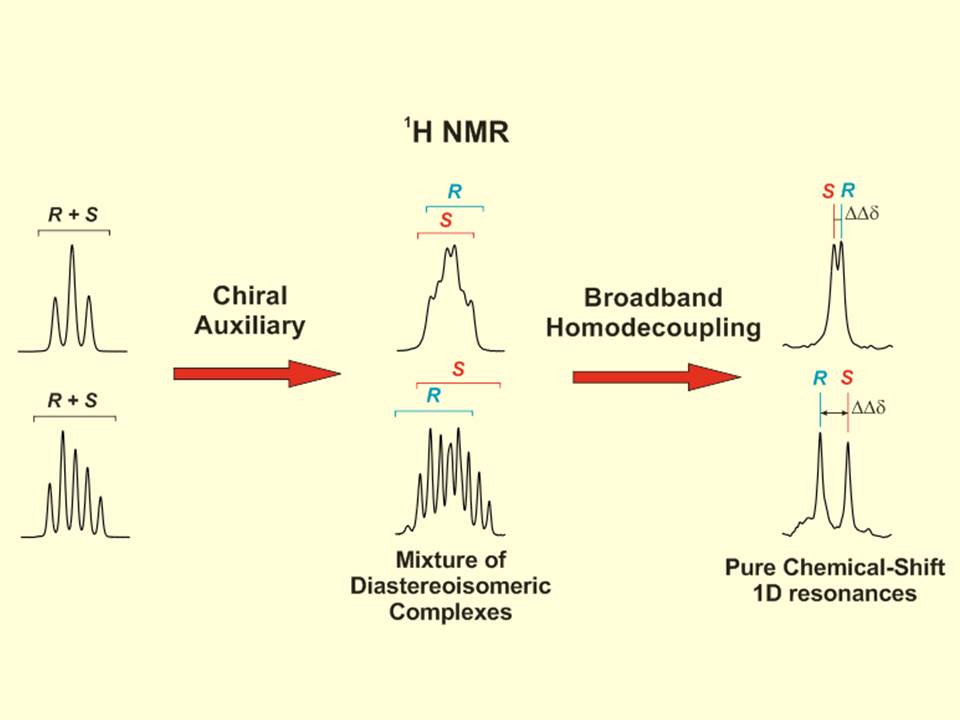
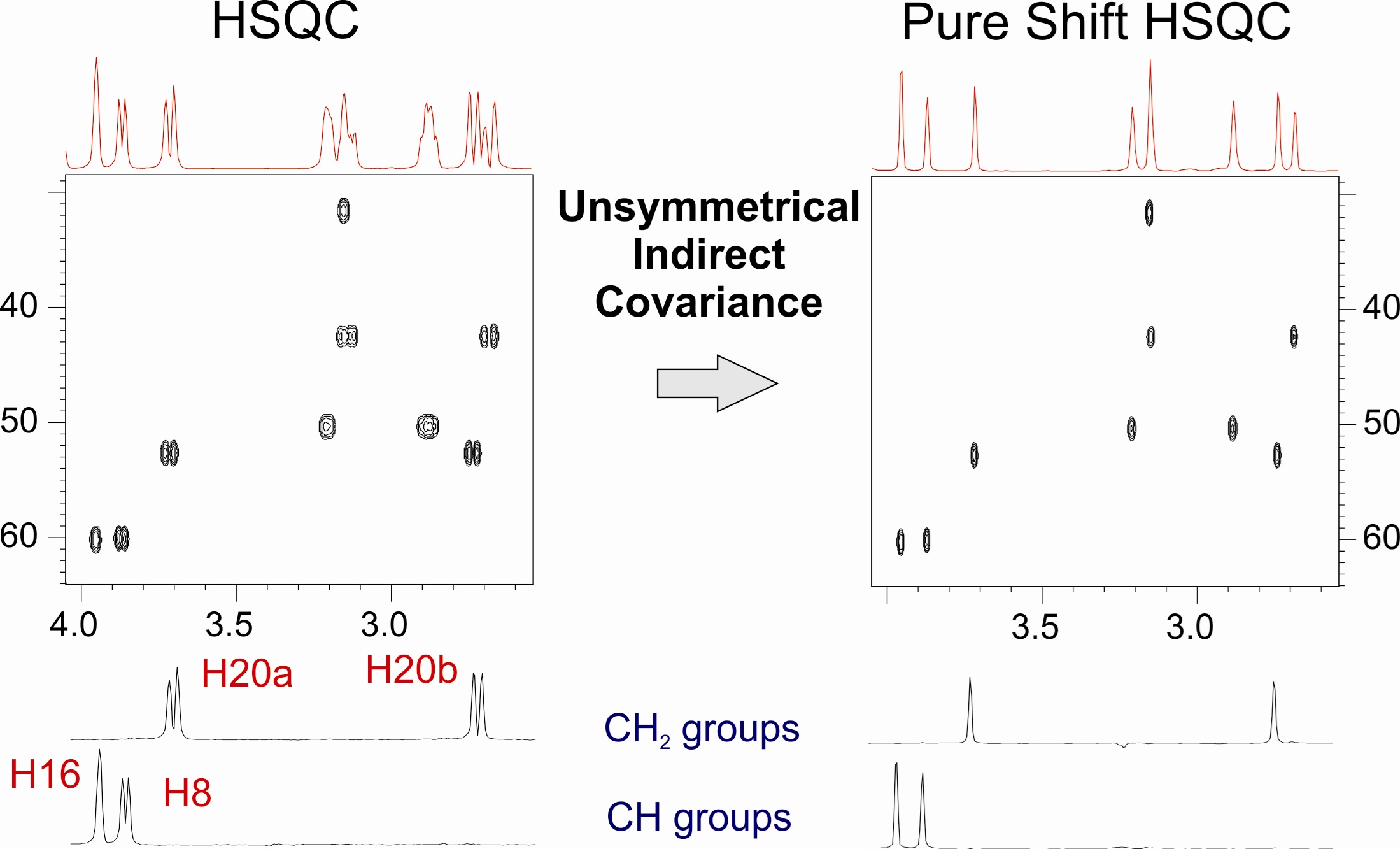
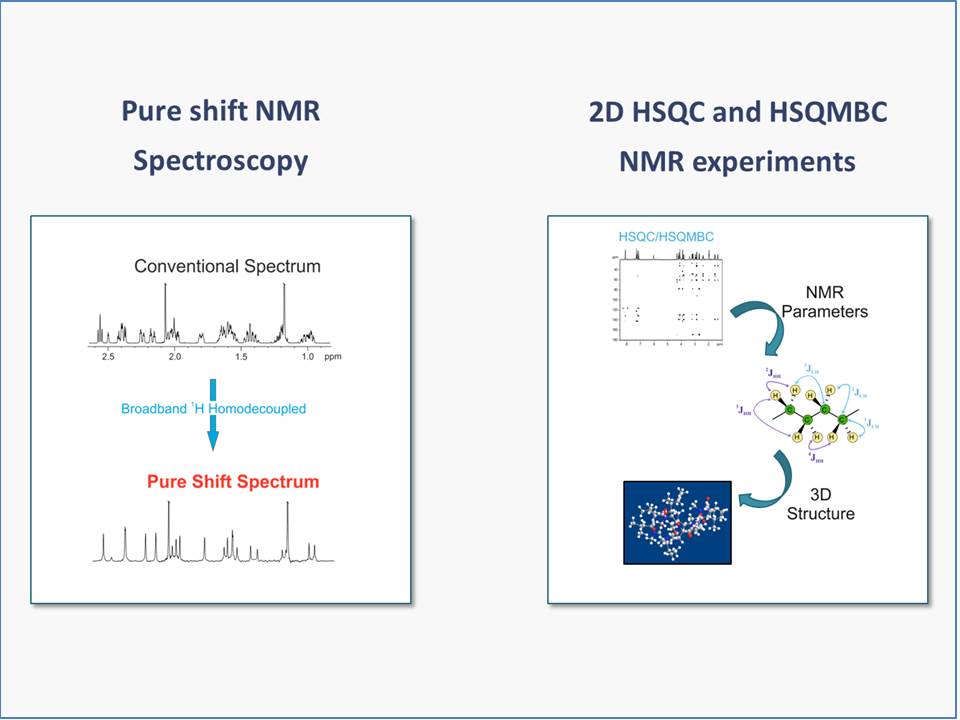
NMR-aided discrimination of enantiomers using chiral solvating agents (CSAs) is a well established method of enantiodifferentiation and measurement of enantiomeric ratios (er). The analysis is traditionally performed by observing chemical shift differences (ΔΔδ) in 1H signals by conventional 1D 1H NMR spectra. However, low ΔΔδ values and signal overlap caused by complex multiplets lead to the lack of spectral signal dispersion that preclude a straightforward analysis. The alternative of using 13C NMR spectroscopy can be more advantageous, because singlet signals are analyzed, but its routine use is limited by its low sensitivity.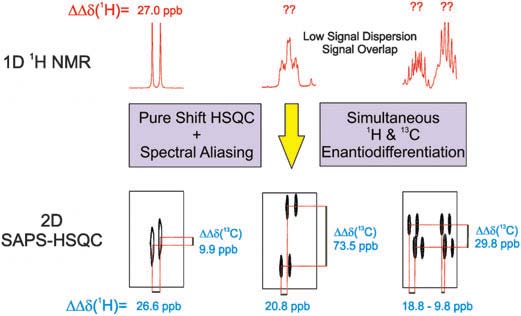 The combination of spectral aliasing and pure shift HSQC in SAPS-HSQC experiments represents an excellent routine tool for NMR enantiodifferentiation studies, yielding simultaneous 1H and 13C enantiodifferentiated data in short times and with high digital resolution and signal dispersion for both nuclei. Its use increases significantly the probability to detect an enantiodifferentiated nucleus, overlapping problems of common 1D 1H experiments are overcome, and poor enantiodifferentiation in 1D experiments can now be detected. The method is compatible with other heteronuclei, with the use of other chiral auxiliaries, and it can be of special interest for enantiodifferentiation experiments of complex mixtures.
The combination of spectral aliasing and pure shift HSQC in SAPS-HSQC experiments represents an excellent routine tool for NMR enantiodifferentiation studies, yielding simultaneous 1H and 13C enantiodifferentiated data in short times and with high digital resolution and signal dispersion for both nuclei. Its use increases significantly the probability to detect an enantiodifferentiated nucleus, overlapping problems of common 1D 1H experiments are overcome, and poor enantiodifferentiation in 1D experiments can now be detected. The method is compatible with other heteronuclei, with the use of other chiral auxiliaries, and it can be of special interest for enantiodifferentiation experiments of complex mixtures.
Pulse Program Code for Bruker: