Some of the SeRMN staff presented our last research works at the annual meeting of the European magnetic resonance community EUROMAR 2017 Conference that took place from 2th to 6th July in Warsaw, Poland. Find below a summary of our contributions.
Teodor Parella presented a lecture entitled: “Accurate measurement of homonuclear and heteronuclear coupling constants from highly resolved HSQC spectra”
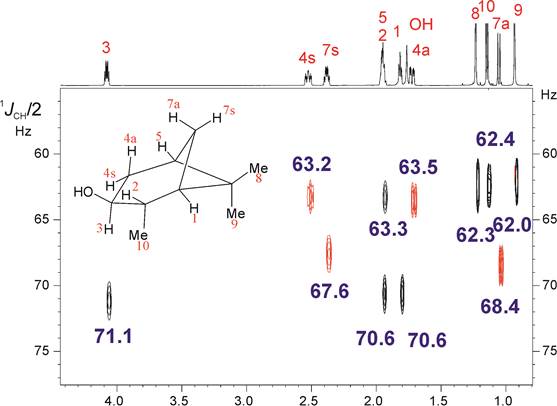
Abstract: The HSQC experiment is a fundamental NMR tool for the structural elucidation of small molecules in solution. In addition, of its conventional implementation to correlated 1H and 13C chemical shifts, it is the basis for a number of powerful solutions such as the measurement of coupling constants. We describe here several approaches to enhance the spectral and digital resolution in a family of HSQC experiments. The incorporation of some NMR concepts such as pure shift NMR, perfect NMR or spectral aliasing will be discussed. In addition, a complete family of J-resolved HSQC pulse schemes will be presented for the accurate determination of both the sign and the magnitude of a different number of homonuclear and heteronuclear coupling constants. A discussion about the advantages to measure them from the indirect F1 dimension will be made. Finally, the application of these novel methods to the measurement of residual dipolar couplings (RDCs) and the role of the concerted used of 1DCH/2DHH for efficient structural discrimination of small molecules will be given.
Pau Nolis presented a poster entitled: “Accurate measurement of JHH in overlapped signals by a TOCSY‐edited SERF Experiment”. André Fredi, Pau Nolis and Teodor Parella.
Abstract: Selective refocusing (GSERF or the recent PSYCHEDELIC) experiments were originally designed to determine all proton–proton coupling constants (JHH) for a selected proton resonance. They work for isolated signals on which selective excitation can be successfully applied but, as it happens in other selective experiments, fail for overlapped signals. To circumvent this limitation, a doubly-selective TOCSY-GSERF scheme is presented for the measurement of JHH in protons resonating in crowded regions. This new experiment takes advantage of the editing features of an initial TOCSY transfer to uncover hidden resonances that become accessible to perform the subsequent frequency-selective refocusing.
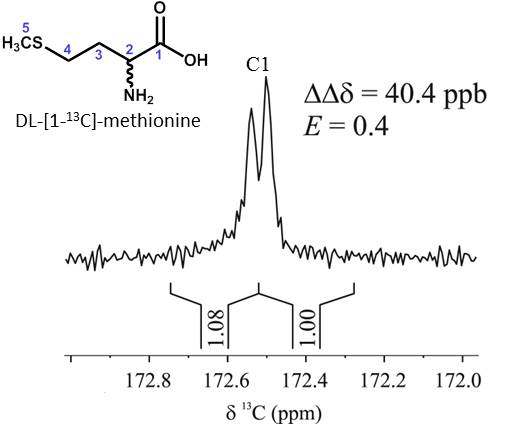
Eva Monteagudo presented a poster entitled: “Chiral recognition by dissolution DNP 13C NMR spectroscopy of DL-[1-13C]methionine”. Eva Monteagudo, Albert Virgili, Teodor Parella and Míriam Pérez-Trujillo. DOI: 10.1021/acs.analchem.7b00156
Abstract: A method based on dissolution DNP 13C NMR spectroscopy to study chiral recognition is described for the first time. 1H NMR experiments using a chiral solvating agent (CSA) as chiral auxiliary are usually performed to measure the enantiomeric excess (ee) of chiral mixtures. However, both overlapping and multiplicity of signals often hamper the enantiomeric differentiation. These drawbacks can be avoided using proton decoupled 13C NMR spectroscopy due to the simplicity of the obtained signals (singlets) and its larger chemical shift range. Unfortunately, the intrinsic low natural abundance of 13C nuclei (1.1 %) means poor sensitivity, thus often long acquisition times. Dissolution DNP (d-DNP) offers a smart solution to sensitivity problems increasing signal enhancements of several orders of magnitude Here in we show how hyperpolarized DL-[1-13C]-methionine enantiomers were differently observed with a single-scan 13C NMR experiment in a millimolar aqueous solution using (-)-(18-crown-6)-2,3,11,12-tetracarboxylic acid as CSA. The developed method entails a step forward in the chiral recognition of small molecules by NMR spectroscopy, its advantages and current limitations, as well as future perspectives are showed and briefly discussed.
Lars Khun presented a poster entitled: “NMR-aided differentiation of enantiomers: Signal enantioresolution”. Míriam Pérez-Trujillo, Teodor Parella and Lars T. Kuhn. DOI: 10.1016/j.aca.2015.02.069
Abstract: The NMR-aided enantiodiscrimination of chiral compounds using chiral auxiliaries (CA) is a widely established method for the differentiation of enantiomers and for measuring enantiomeric ratios (er). Up to the present, the study, optimization, and comparison of such methods have been performed based on the enantiodifferentiation of NMR signals via analyzing non-equivalent chemical shift values (ΔΔδ) of the diastereoisomeric species formed. In the large majority of cases, however, a poor and non-reliable comparison of results is often obtained via the analysis of ΔΔδ exclusively.
In here, the concept of enantioresolution of an individual NMR signal and its importance for NMR-aided enantiodifferentiation studies is introduced and discussed. In addition, the enantioresolution quotient, E, is proposed as the parameter to describe its quantification. Using three structurally very different biochemically active molecules (tryptophan, 1-aminoindan, ibuprofen), we show that, in comparison to measuring ΔΔδ, the experimental determination of E allows a more reliable interpretation of the results stemming from NMR-aided enantiodiscrimination experiments and opens up new possibilities for the study of enantiodifferentiation data derived from novel NMR experiments, setup improvements or new CAs. We envisage that useful applications involving the application of E will comprise the analysis of enantiodifferentiated signals derived from (i) different NMR experiments, e.g., 1D 1H NMR standard vs. 1H pure shift experiments, as the use of E will consider the differences in the multiplet pattern of an enantiodifferentiated signal; (ii) using different nuclei and NMR experiments, e.g., 1D 1H vs. 1D 13C NMR signals from standard experiments; (iii) methodological improvements, e.g., hyperpolarization methods; and (iv) new CSAs as well as prochiral solvating agents (pro-CSAs).
Save